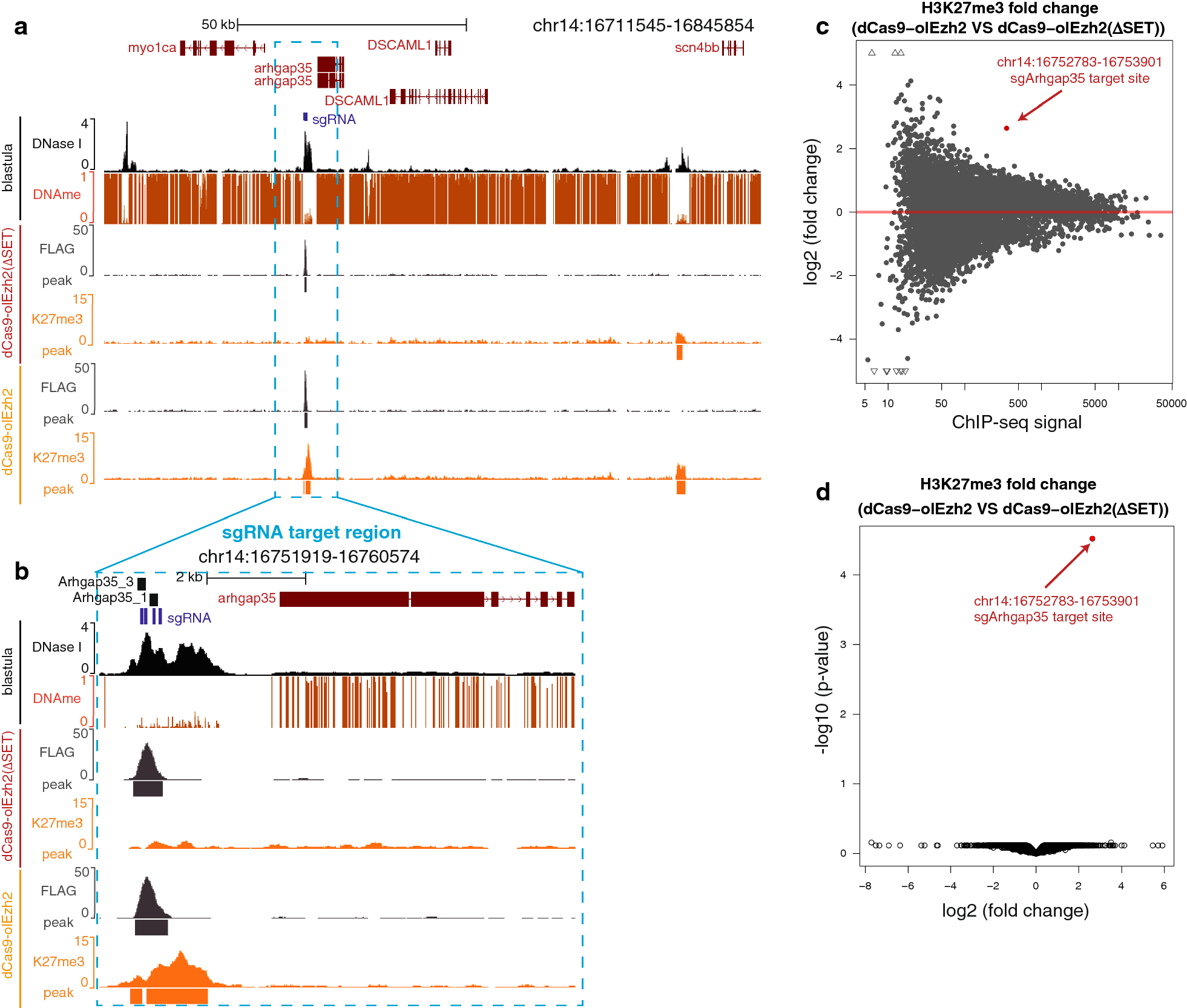
Validation of small-scale ChIP-seq histone mark maps. (A) Two H3K27me3... | Download Scientific Diagram

Local difference of H3K27me3 marks in mutants of PcG components. (A)... | Download Scientific Diagram
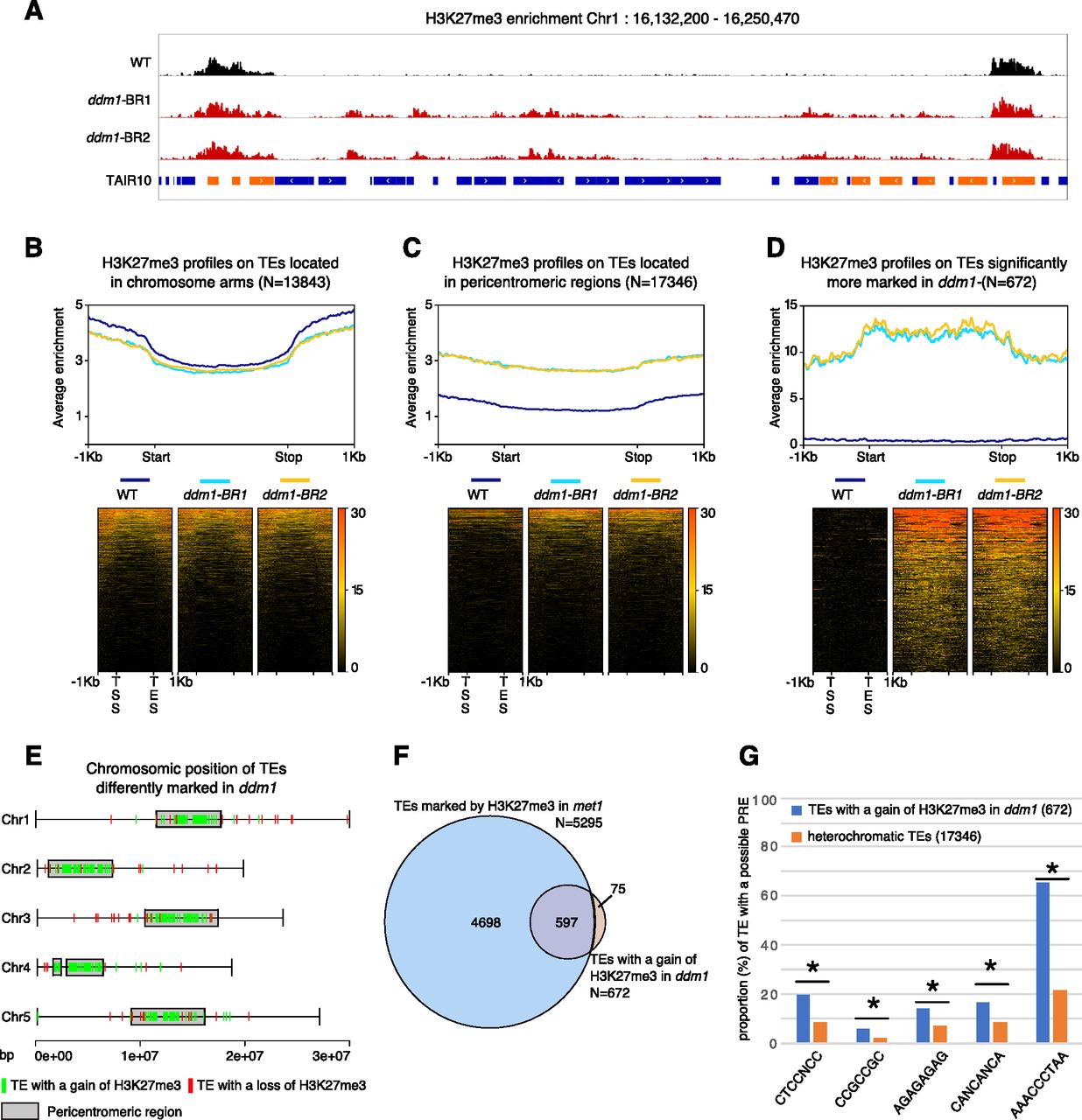
Polycomb mutant partially suppresses DNA hypomethylation–associated phenotypes in Arabidopsis | Life Science Alliance

Quantitative Multiplexed ChIP Reveals Global Alterations that Shape Promoter Bivalency in Ground State Embryonic Stem Cells - ScienceDirect

Global changes of H3K27me3 domains and Polycomb group protein distribution in the absence of recruiters Spps or Pho | PNAS

Genome‐wide occupancy of histone H3K27 methyltransferases CURLY LEAF and SWINGER in Arabidopsis seedlings - Shu - 2019 - Plant Direct - Wiley Online Library

Genome-wide localization patterns by ChIP-seq of H3K27me3, Su(z)12, and... | Download Scientific Diagram
Genomewide Analysis of PRC1 and PRC2 Occupancy Identifies Two Classes of Bivalent Domains | PLOS Genetics
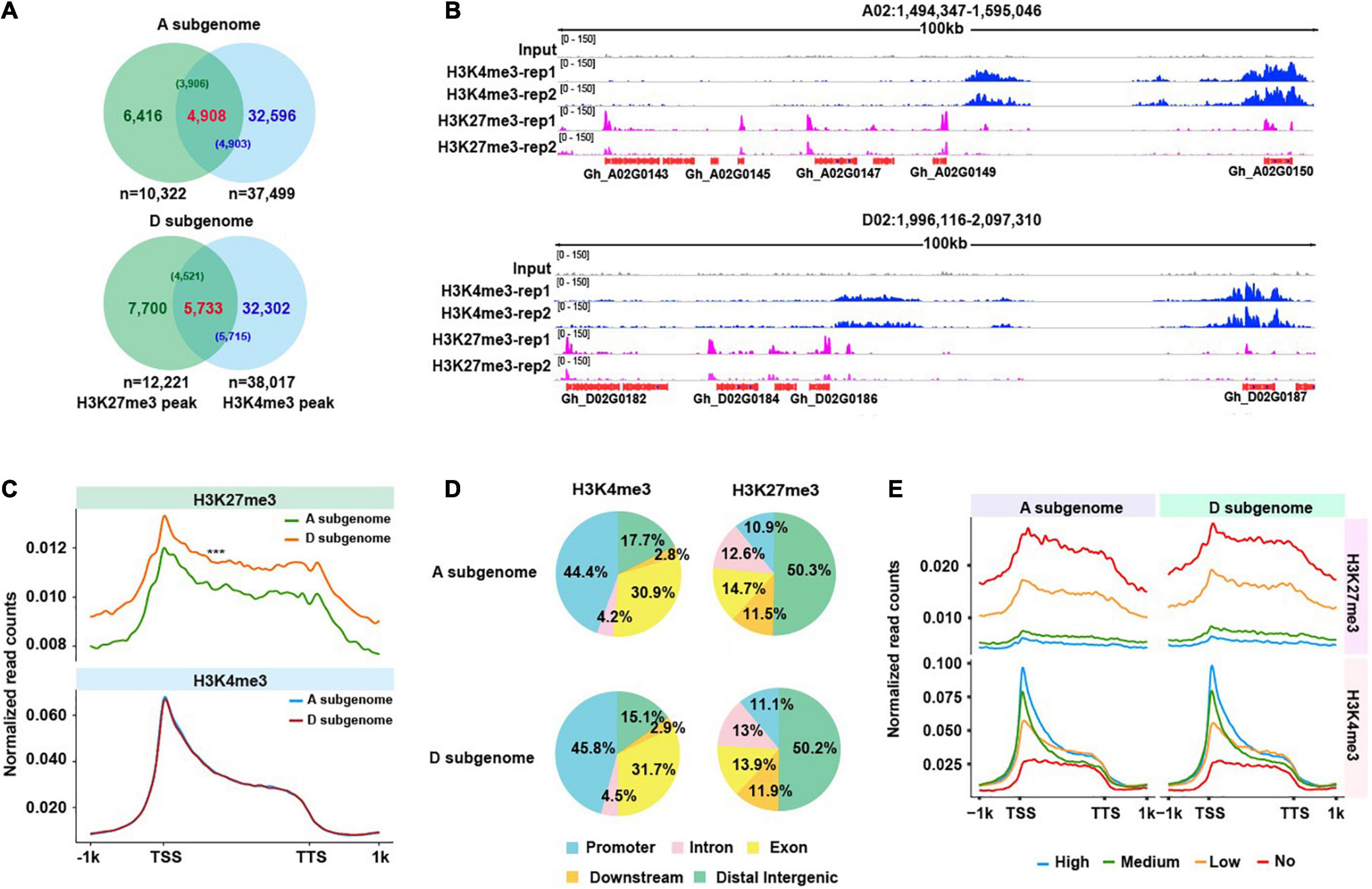
Frontiers | Profiling of H3K4me3 and H3K27me3 and Their Roles in Gene Subfunctionalization in Allotetraploid Cotton

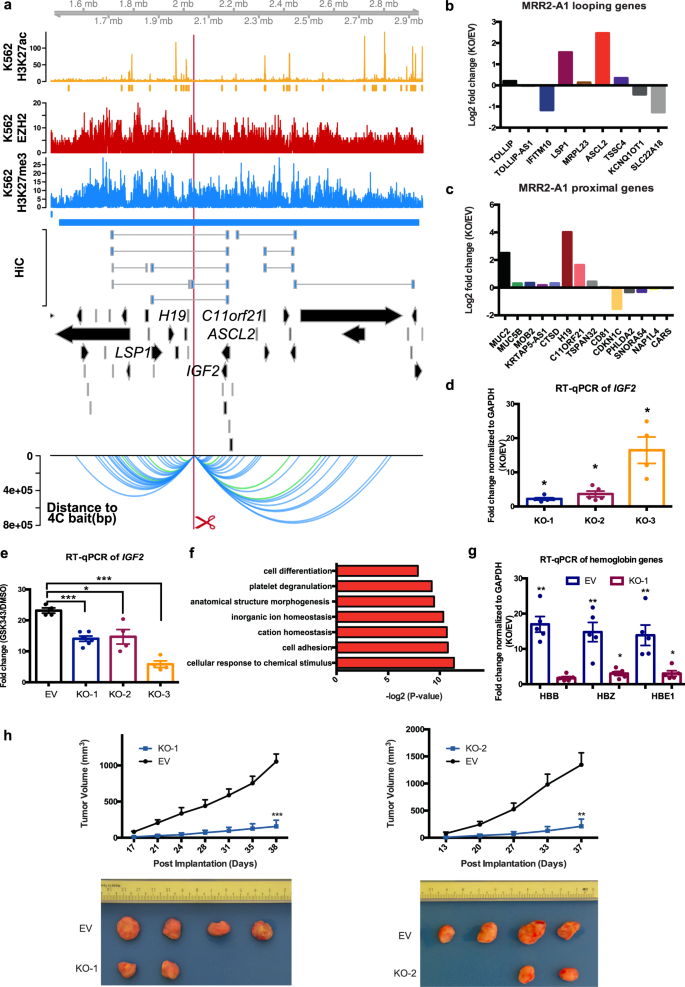
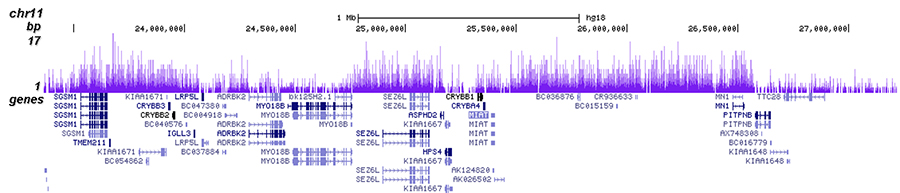
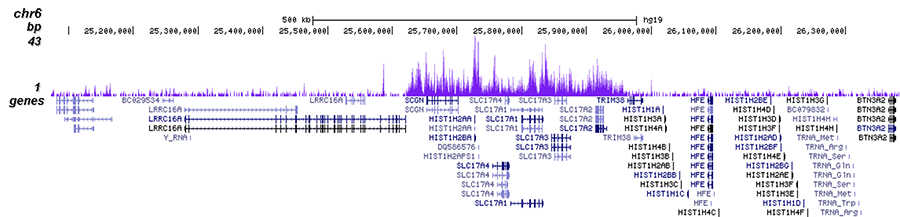
H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions | bioRxiv
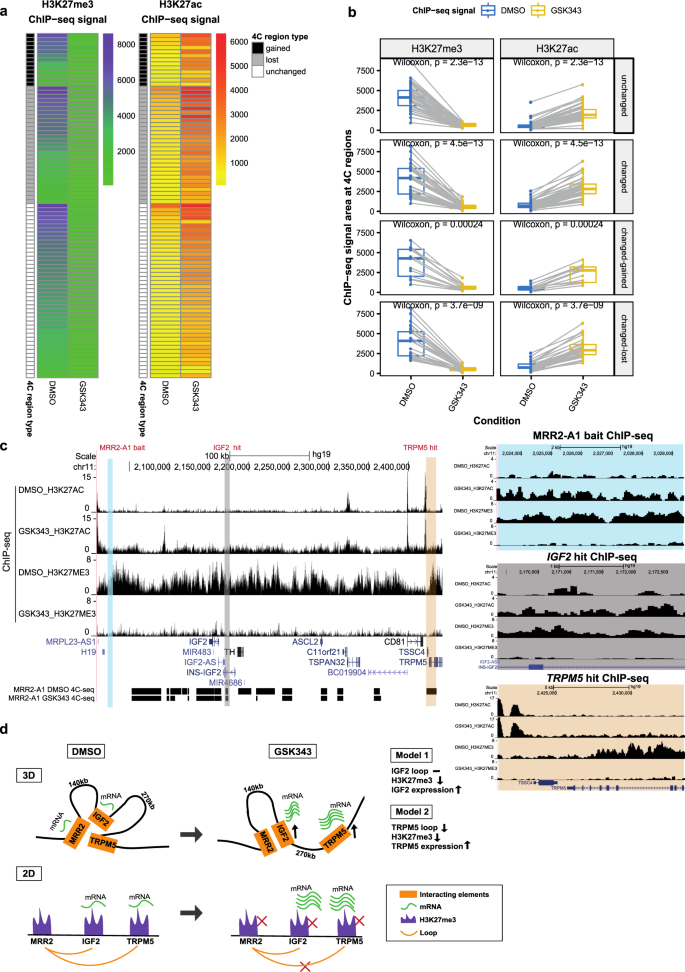
H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions | Nature Communications

Global changes of H3K27me3 domains and Polycomb group protein distribution in the absence of recruiters Spps or Pho | PNAS
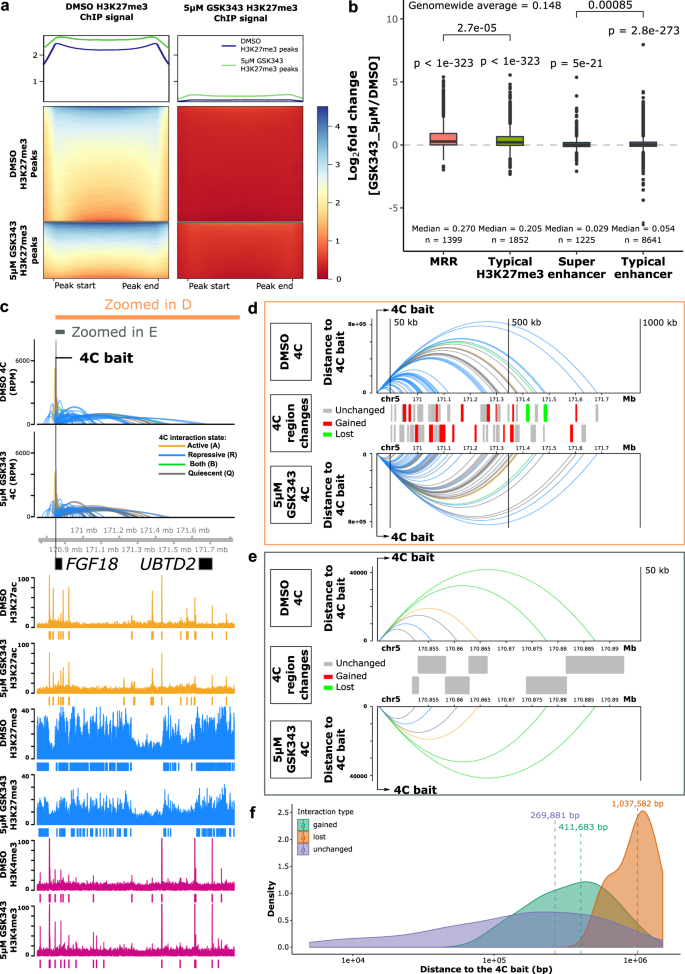
H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions | Nature Communications

H4K20me1 and H3K27me3 are concurrently loaded onto the inactive X chromosome but dispensable for inducing gene silencing | EMBO reports
Genomewide Analysis of PRC1 and PRC2 Occupancy Identifies Two Classes of Bivalent Domains | PLOS Genetics
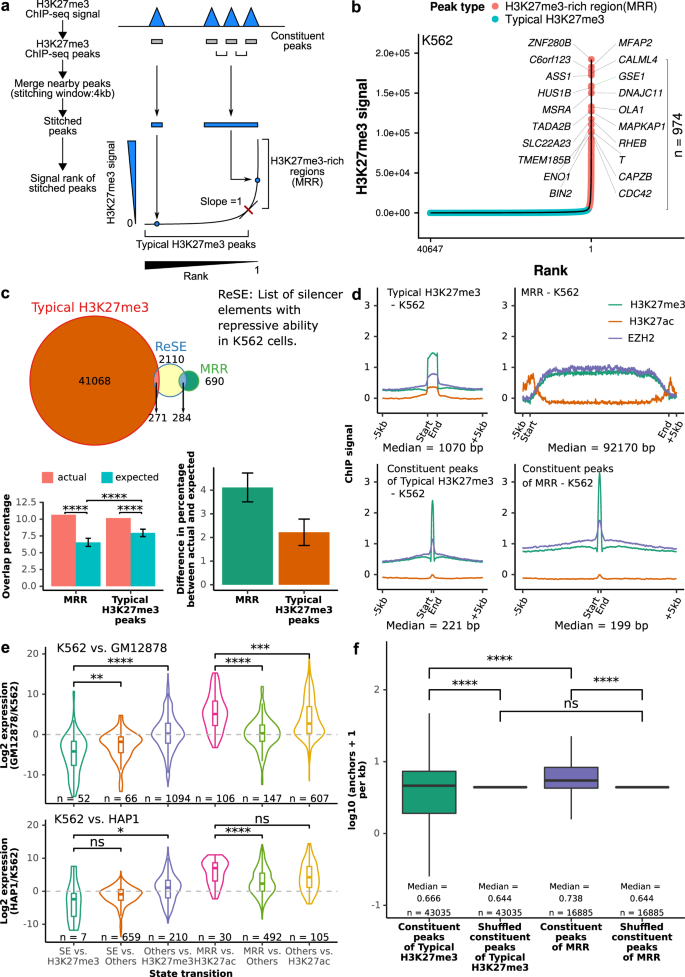
H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions | Nature Communications
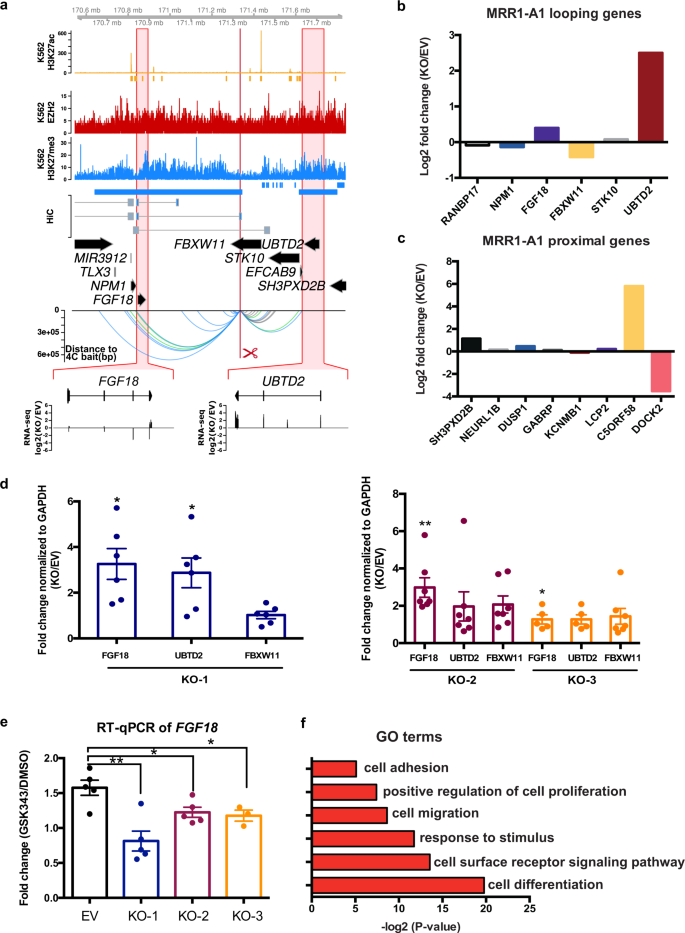
H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions | Nature Communications
Nucleosome Density ChIP-Seq Identifies Distinct Chromatin Modification Signatures Associated with MNase Accessibility
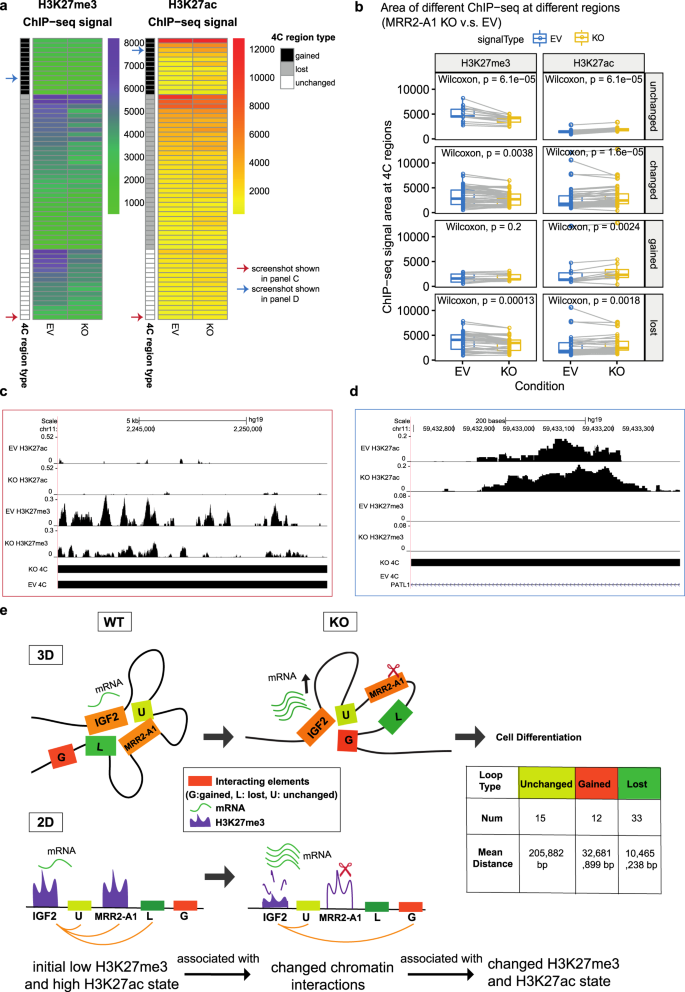
H3K27me3-rich genomic regions can function as silencers to repress gene expression via chromatin interactions | Nature Communications
![PDF] ChIP-seq analysis reveals distinct H3K27me3 profiles that correlate with transcriptional activity | Semantic Scholar PDF] ChIP-seq analysis reveals distinct H3K27me3 profiles that correlate with transcriptional activity | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/401cd71658c42f6b87c32dc57a34050ed6929121/8-Figure4-1.png)
PDF] ChIP-seq analysis reveals distinct H3K27me3 profiles that correlate with transcriptional activity | Semantic Scholar

ChIP-seq of EZH2 and H3K27me3 in WT iPSCs a, Comparison of the EZH2... | Download Scientific Diagram